Visualisation Functions (Python)
This page lists the Statescope visualisation helpers and when to use them. Most plots expect that you have run the relevant module (Deconvolution, Refinement, or StateDiscovery) first.
Quick start
from Statescope import Initialize_Statescope
from Statescope import (
Heatmap_Fractions,
Boxplot_CelltypeAbundance,
Heatmap_GEX,
Heatmap_StateScores,
Heatmap_StateLoadings,
Plot_CopheneticCoefficients,
BarPlot_StateLoadings,
TSNE_AllStates,
TSNE_CellTypes,
)
Statescope_model = Initialize_Statescope(Bulk, TumorType="PBMC", Ncelltypes=7, Ncores=10)
Statescope_model.Deconvolution()
Statescope_model.Refinement()
Statescope_model.StateDiscovery()
Deconvolution outputs
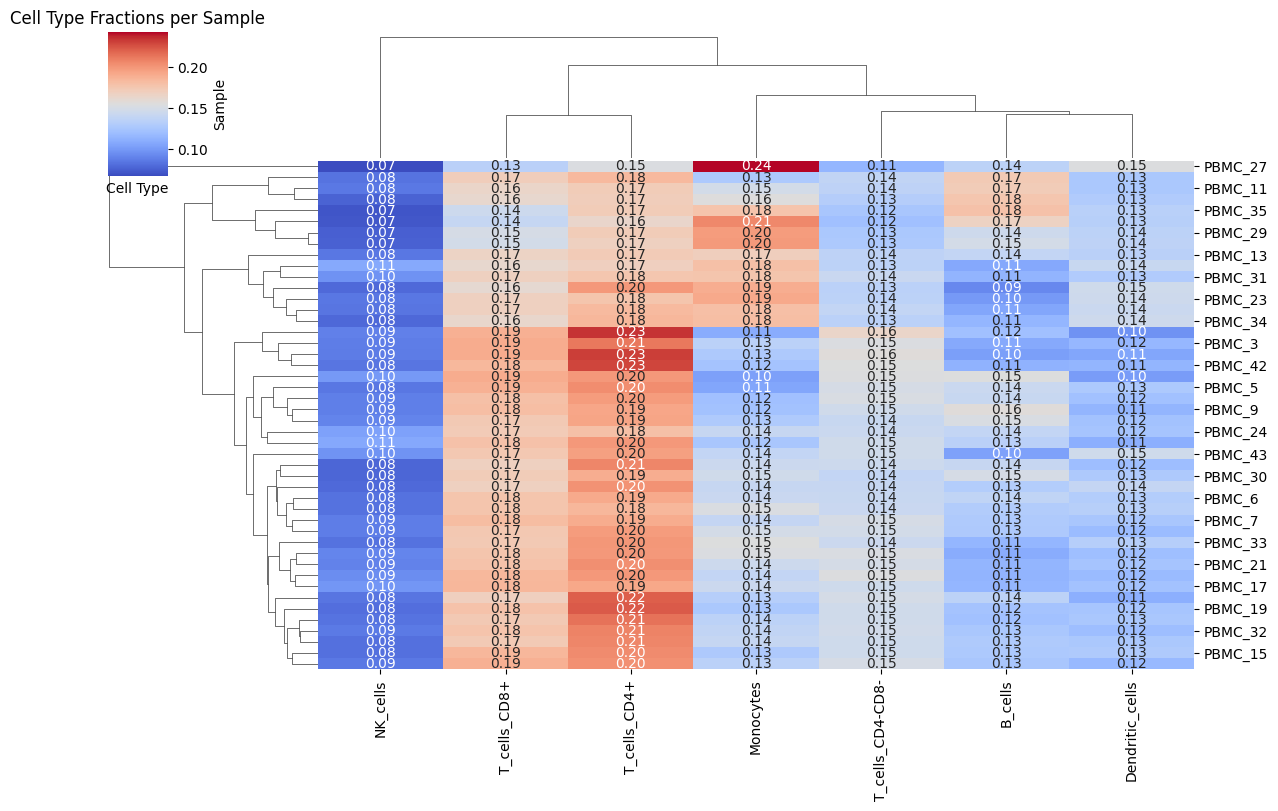
Heatmap_Fractions
Visualizes cell-type fractions across samples.
Heatmap_Fractions(Statescope_model)
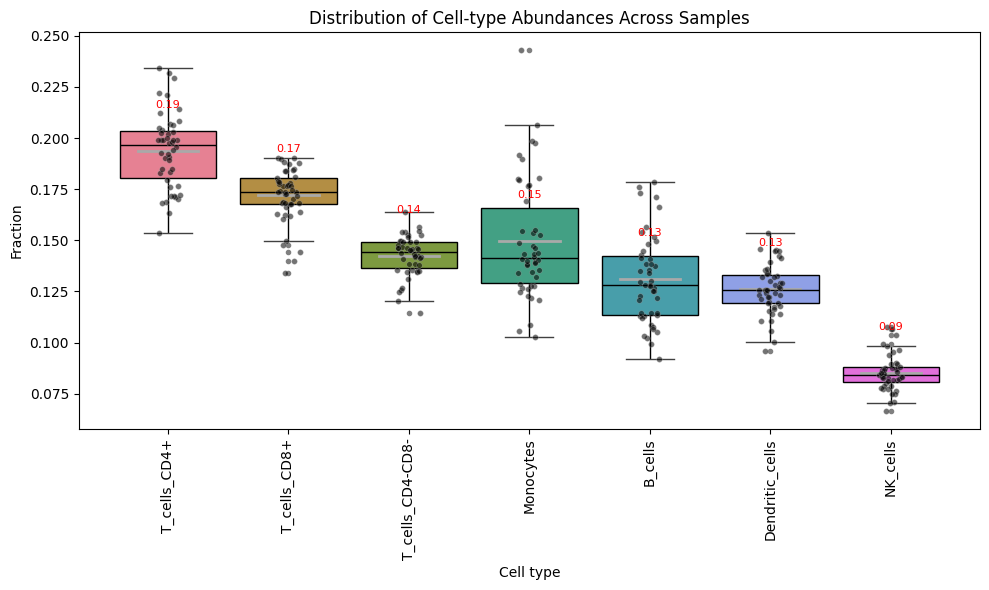
Boxplot_CelltypeAbundance
Boxplot of cell-type fractions across samples.
Boxplot_CelltypeAbundance(Statescope_model)
Refinement outputs
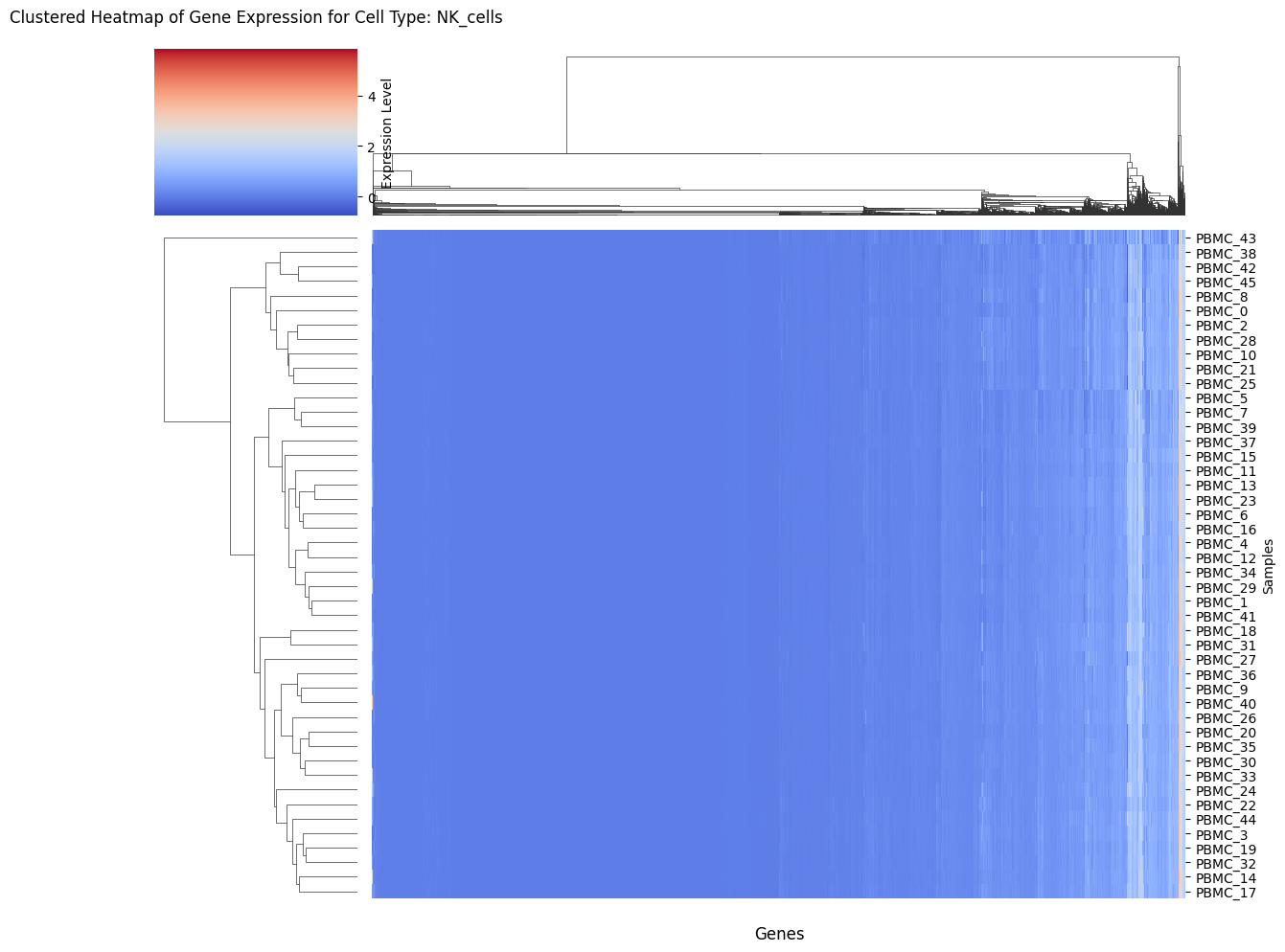
Heatmap_GEX
Clustered heatmap of purified gene expression for a cell type.
Heatmap_GEX(Statescope_model, "T_cells_CD4+")
State discovery outputs
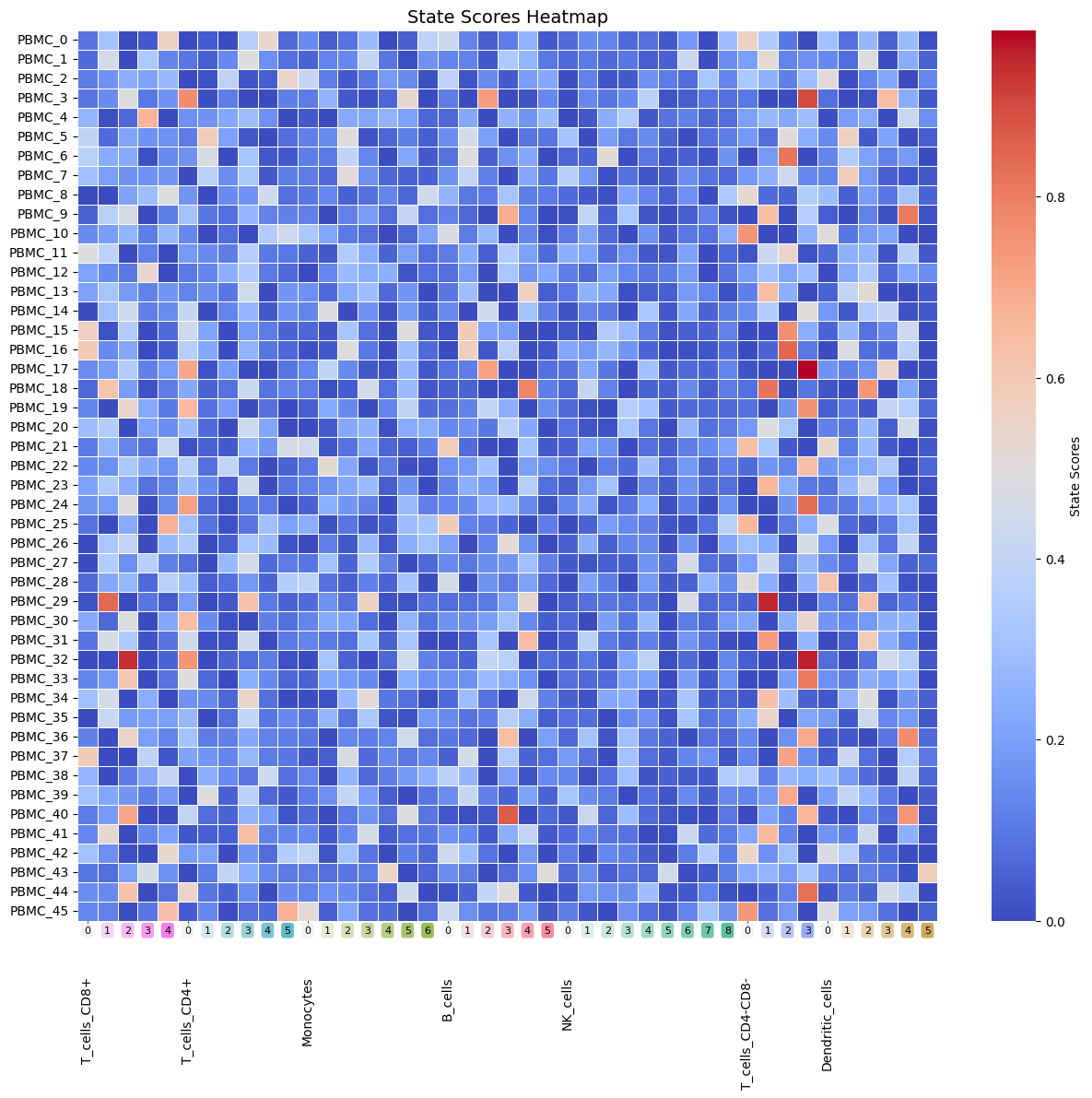
Heatmap_StateScores
Heatmap of per-sample state scores.
Heatmap_StateScores(Statescope_model)
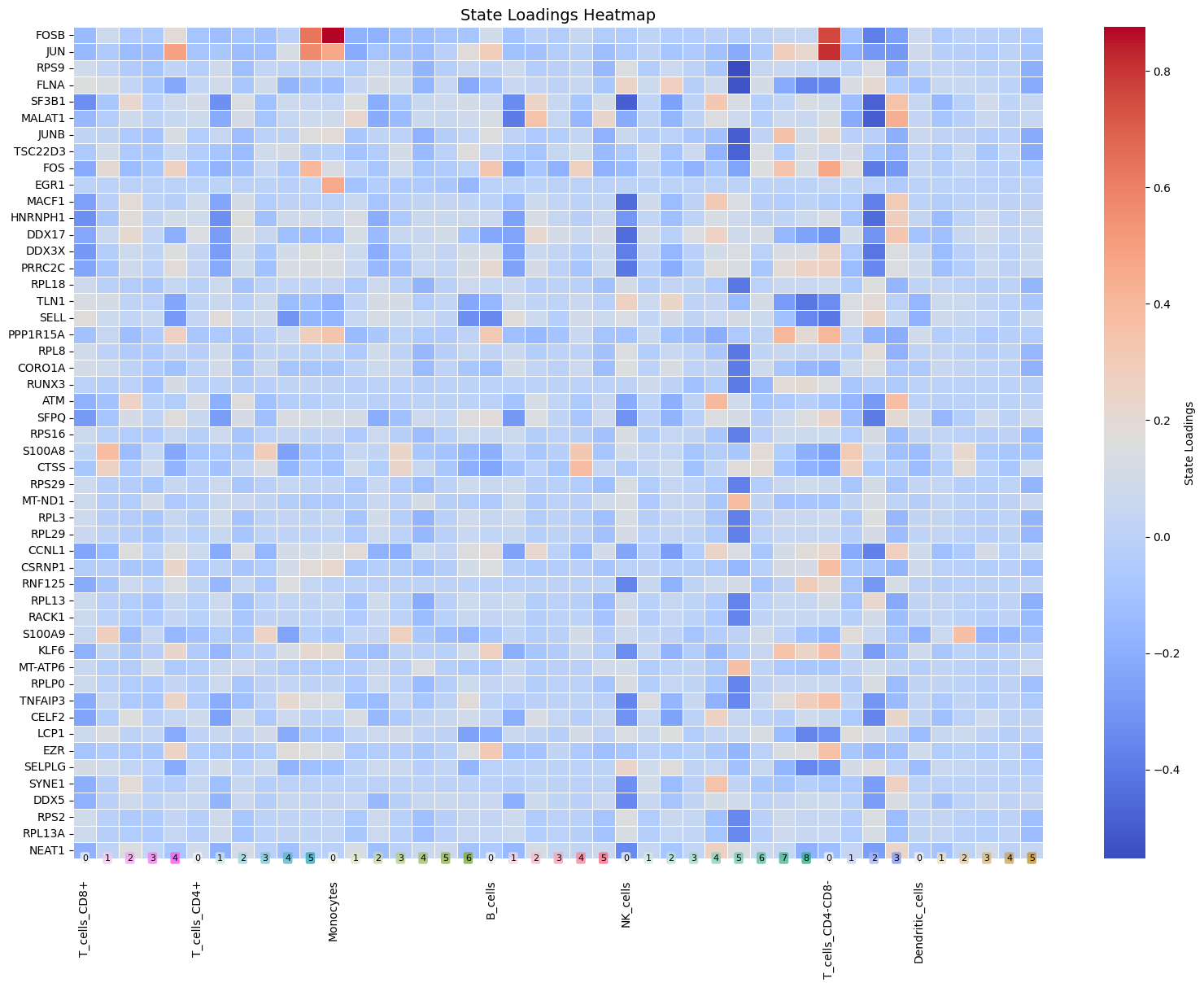
Heatmap_StateLoadings
Heatmap of state loadings; use top_genes to reduce clutter.
Heatmap_StateLoadings(Statescope_model, top_genes=50)
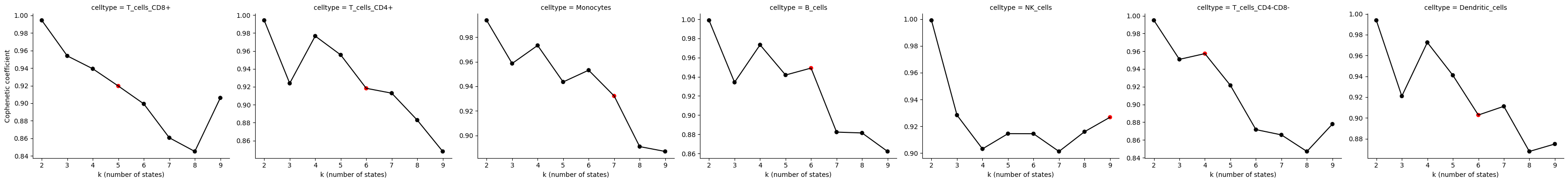
Plot_CopheneticCoefficients
Plots cophenetic coefficients across k values (used for model selection).
Plot_CopheneticCoefficients(Statescope_model)
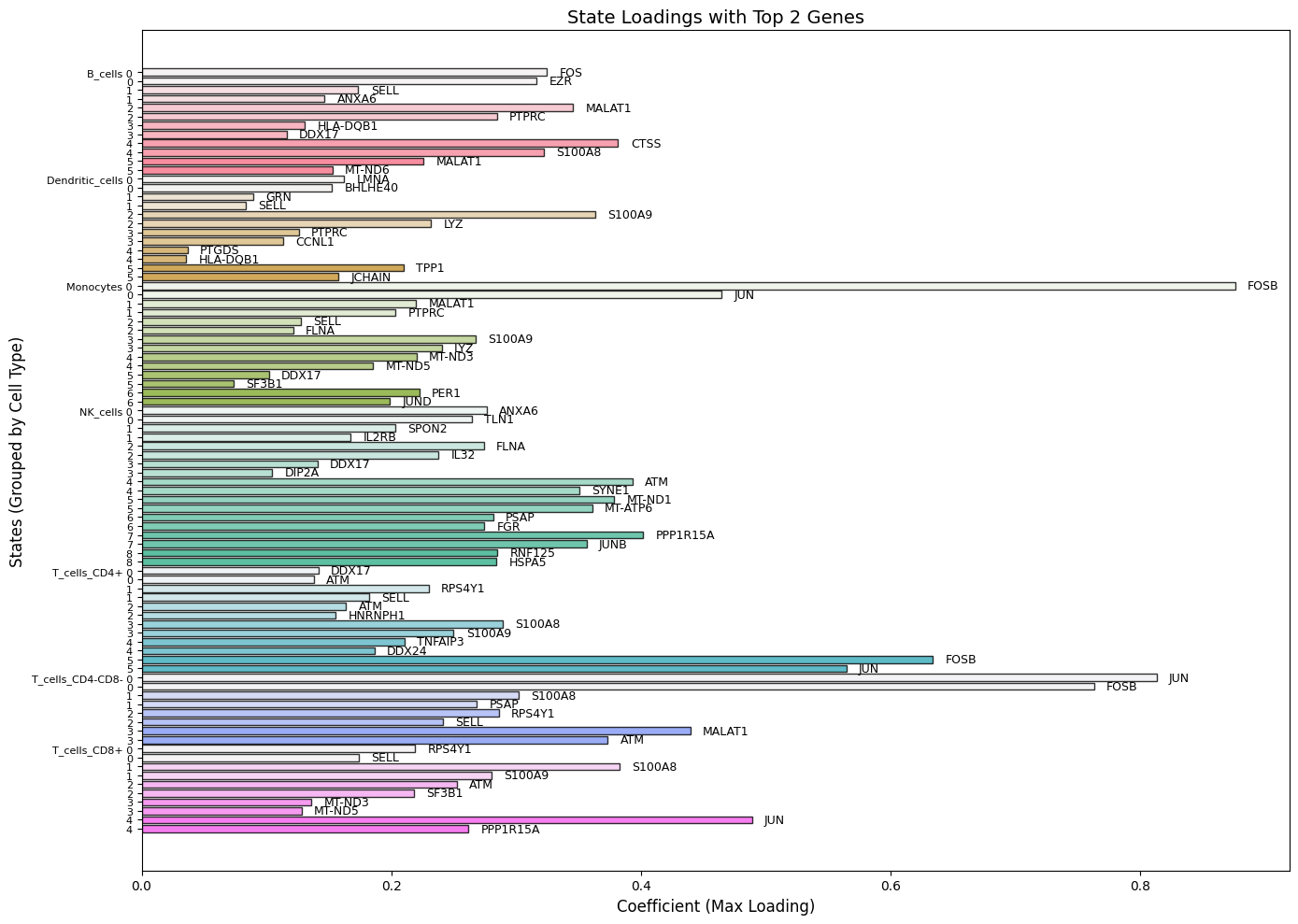
BarPlot_StateLoadings
Bar plot of state loadings with top genes labeled.
BarPlot_StateLoadings(Statescope_model, top_genes=3)
TSNE_AllStates
t-SNE for all cell types and states (optionally show sample names).
TSNE_AllStates(Statescope_model, show_samples=True)
TSNE_CellTypes
t-SNE for one or more cell types.
TSNE_CellTypes(Statescope_model, celltype="Epithelial", show_samples=True)
Usage notes
- Run
Deconvolution()before fraction plots. - Run
Refinement()before GEX heatmaps. - Run
StateDiscovery()before state score/loading plots and t-SNE views. - It is recommended to save your model after each module so you can reload and continue analysis later.